Note
Go to the end to download the full example code.
Orientation from aligning directions#
This example demonstrates how to use
from_align_vectors() to estimate an orientation
\(O\) from two sets of aligned vectors.
One set of vectors \(\mathbf{v}\) is given in the sample reference reference frame,
\((x, y, z)\), the other set \(\mathbf{t}\) is given in the crystal reference
frame, \((e_1, e_2, e_3)\).
Vector angle deviation [deg]: 3.2979238394332837
Error distance: 0.01642441765779543
from diffpy.structure import Lattice, Structure
import matplotlib.pyplot as plt
import numpy as np
from orix.crystal_map import Phase
from orix.quaternion import Orientation
from orix.vector import Miller, Vector3d
plt.rcParams.update({"figure.figsize": (5, 5), "lines.markersize": 8})
# Specify an hexagonal crystal structure and symmetry
phase = Phase(
point_group="6/mmm",
structure=Structure(lattice=Lattice(1, 1, 2, 90, 90, 120)),
)
# Define a reference orientation (goal)
O_ref = Orientation.from_axes_angles([1, 1, 1], -45, phase.point_group, degrees=True)
# Specify two crystal directions (any will do)
t = Miller(uvw=[[2, 1, 1], [1, 3, 1]], phase=phase)
# Find out where these directions in the reference orientation (crystal)
# point in the sample reference frame
v = Vector3d(~O_ref * t)
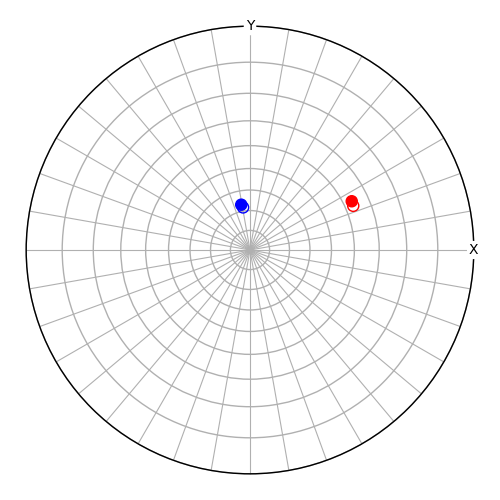
# Plot the reference orientation sample directions as empty circles
fig = v.scatter(
c="none",
ec=["r", "b"],
grid=True,
axes_labels=["X", "Y"],
return_figure=True,
figure_kwargs={"layout": "tight", "figsize": (5, 5)},
)
# Add some randomness to the sample directions (0 error magnitude gives
# exact result)
err_magnitude = 0.1
v_err = v + Vector3d(np.random.normal(0, err_magnitude, 3))
angle_err = v_err.angle_with(v, degrees=True).mean()
print("Vector angle deviation [deg]: ", angle_err)
# Obtain the orientation which aligns the crystal directions with the
# sample directions
O_new, err = Orientation.from_align_vectors(t, v_err, return_rmsd=True)
print("Error distance: ", err)
# Plot the crystal directions in the new orientation
v2 = Vector3d(~O_new * t)
v2.scatter(c=["r", "b"], figure=fig)
Total running time of the script: (0 minutes 0.277 seconds)
Estimated memory usage: 493 MB